 |
 |
- Search
| Genomics Inform > Volume 19(4); 2021 > Article |
|
Abstract
Predicting individual traits and diseases from genetic variants is critical to fulfilling the promise of personalized medicine. The genetic variants from genome-wide association studies (GWAS), including variants well below GWAS significance, can be aggregated into highly significant predictions across a wide range of complex traits and diseases. The recent arrival of large-sample public biobanks enables highly accurate polygenic predictions based on genetic variants across the whole genome. Various statistical methodologies and diverse computational tools have been introduced and developed to compute the polygenic risk score (PRS) more accurately. However, many researchers utilize PRS tools without a thorough understanding of the underlying model and how to specify the parameters for the best performance. It is advantageous to study the statistical models implemented in computational tools for PRS estimation and the formulas of parameters to be specified. Here, we review a variety of recent statistical methodologies and computational tools for PRS computation.
Accurately predicting complex traits and diseases (e.g., type 2 diabetes, cancer, and asthma) based on an individual’s genetic variants is crucial for effective disease prevention and personalized treatment [1-8]. The genetic architecture of many diseases contains a substantial polygenic component, meaning that thousands of variants with small effects contribute to disease risk. This limited the predictive ability of genetic variants in early studies based on significant associations from genome-wide association studies (GWAS). However, genetic variants—mostly single-nucleotide polymorphisms (SNPs)—from GWAS, including variants well below genome-wide significance, can be aggregated into highly significant predictions of phenotypes across a wide range of complex traits and diseases. With the recent arrival of public biobanks with 500K‒1M samples, highly accurate polygenic prediction is poised to become a reality [9]. The emergence of highly accurate polygenic prediction has led to the revitalization of the polygenic risk score (PRS), which is the score for predicting a trait and disease risk, calculated as the weighted sum of risk alleles with predicted weights computed by coefficients from GWAS.
For accurate PRS estimation, various statistical methodologies have been proposed and diverse computational tools have been developed, such as PLINK (https://zzz.bwh.harvard.edu/plink/) [10], GCTA (Genome-wide Complex Trait Analysis, https://cnsgenomics.com/software/gcta/) [11], and LDpred (https://github.com/bvilhjal/ldpred) [12]. These PRS tools have been widely adopted for genetic risk prediction in practice due to their easy usability with a proper theoretical basis. These tools compute the PRS using various data types, including the individual-level genotype as well as summary-level data on the basis of multiple regression, linear mixed models (LMMs), and Bayesian models. Despite the popularity of PRS tools, many researchers have utilized them without a thorough understanding of the underlying model and how to specify the parameters for the best performance. To achieve better prediction, it is advantageous to study the statistical models implemented in computational tools for PRS estimation and the mathematical formula of parameters to be specified. A deep understanding of the underlying statistical models in PRS software and a comparison of their advantages and disadvantages will help researchers to find an appropriate PRS tool for themselves.
Here, we review various statistical methodologies and computational tools for PRS computation. First, we review summary-based PRS methods with a few published SNPs or whole SNPs from large-sample GWAS using PLINK [10] and LDpred [12], with two main considerations: non-infinitesimal genetic architecture and the linkage disequilibrium (LD) structure of the genotype data. Second, we review traditional best linear unbiased prediction (BLUP)-based prediction with individual-level genotypes using genomic BLUP (GBLUP) [13] and summary-level data using summary statistics BLUP (SBLUP) [14]. Third, we review Bayesian multiple regression (BMR)-based prediction with individual-level data using BayesR [15,16] and summary-level data using summary statistics BayesR (SBayesR) [17]. Fourth, we review penalized regression-based approaches using the least absolute shrinkage and selection operator (lasso) [18,19], the elastic net [20], and lassosum (https://github.com/tshmak/lassosum/) [21]. Fifth, we review statistical methods for jointly analyzing multiple phenotypes to further improve prediction accuracy using multi-trait GBLUP (MTGBLUP, https://github.com/uqrmaie1/mtgblup) [16], weighted multi-trait SBLUP (wMT-SBLUP, https://github.com/uqrmaie1/smtpred) [22], and cross-trait penalized regression (CTPR, http://lianglab.rc.fas.harvard.edu/CTPR/) [23]. Finally, we review multi-ethnic approaches to incorporate information from multiple populations using XP-BLUP (https://github.com/tanglab/XP-BLUP) [24], multi-ethnic PRS [25], and multi-ancestry PRS [26]. We conclude with a discussion of statistical models and computational tools that require further work on improving the accuracy of PRS prediction. Table 1 presents a list of the PRS methods reviewed in this paper, along with their underlying statistical models, computational tools, and required data.
A study of schizophrenia showed that the PRS achieved significantly better prediction in validation samples than random scores, and far more accurate than those based on the single GWAS loci discovered in the study [27]. This study describes an early demonstration of the importance and advantages of the PRS for the prediction of disease risk [28].
Let β ^ j y ^ l = ∑ j β ^ j x i j h G W A S 2 h G W A S 2
The prediction R2 of published SNPs depends on the genetic architecture of the phenotypes. Under an infinitesimal genetic architecture, all SNPs are causal with relatively small effect size, and thus the associated SNPs identified by GWAS studies explain a small amount of genetic variance, achieving poor prediction R2. For example, the narrow-sense heritability (h2) for BMI is h2=0.4-0.6, but the heritability explained by GWAS-significant SNPs with >300K samples yields h G W A S 2 h G W A S 2
Polygenic risk prediction can be performed using all SNPs from GWAS studies, not only GWAS-significant SNPs. The PRS can be estimated as y ^ l = ∑ j β ^ j x i j h g 2 E [ R 2 ] ≈ h g 2 / [ 1 + M / h g 2 N ] h g 2
In order to utilize all SNPs to compute the PRS, there are two main considerations: (1) the non-infinitesimal genetic architecture of the phenotype, and (2) the LD structure of the genotype data. That is, y ^ l = ∑ j β ^ j x i j
Popular genetic tools, such as PLINK [10], PRSice (https://www.prsice.info/) [35], and PRSice-2 [36], are utilized to estimate PRS with a few published SNPs or all SNPs from GWAS studies. The PLINK is not originally designed for PRS computation, but every required procedure of the C+T (LD clumping + p-value thresholding) approach can be performed with PLINK. It requires the summary statistics from GWAS studies as well as phenotype, covariate, and genotype data from target samples after a quality control procedure. To account for the LD structure, LD clumping is performed using the PLINK options (e.g., --clump-r2 0.1 --clump-kb 250) to form clumps of all SNPs that are within a certain distance (in kilobases [kb]) from the index SNPs (e.g., 250 kb) and that are in LD with the index SNP based on the r2 threshold (i.e., r2 < 0.1). For p-value thresholding, the SNPs are generated with p-values less than a provided threshold (PT) and then candidate PRSs corresponding to the thresholds are created with the PLINK options (e.g., --score, --q-score). The best PRS is selected among candidate PRSs computed at a range of p-value thresholds based on the prediction R2. For the automation of the C+T approach in PLINK, we can utilize PRSice and PRSice-2, which are options in R software for computing and evaluating the PRS. PRSice and PRSice-2 are popular PRS tools and constitute efficient and scalable software for automating and simplifying PRS computation on large-scale GWAS data. They handle imputed data as well as genotyped data and simultaneously evaluate a large number of continuous and binary phenotypes. Similar to PLINK, they require summary data as well as phenotype, covariate, and genotype data for the target samples. They automate the procedure of the standard C+T method, which utilizes PLINK options for PRS analysis.
A critical issue in estimating the PRS is the LD structure between SNPs, which has been heuristically addressed by LD pruning and LD clumping. Recently, LDpred was developed as a more sophisticated method that also utilizes summary statistics [12]. It has been shown that modeling LD using an LD reference panel and estimating the posterior mean of effect size can improve prediction accuracy [28].
LDpred is an LD matrix and summary statistics–based Bayesian method for polygenic prediction, which is a popular tool for deriving the PRS [12]. It computes posterior means under a point-normal prior, accounting for LD information. The PRS is computed by y ^ l = ∑ j E ( β j | β ^ j ) x i j y ^ l E ( β j | β ^ j )
In the special case of no LD between SNPs, the posterior mean can be computed analytically. Under a Gaussian infinitesimal prior, β j ~ N ( 0 , h g 2 M ) E ( β j | β ^ j ) = h g 2 h g 2 + M / N β ^ j β ^ j ~ β j + ϵ j , ϵ j ~ N ( 0 , 1 N ) β ^ j β j ~ N ( 0 , h g 2 M p ) E ( β j | β ^ j ) = h g 2 h g 2 + M p / N p ¯ j β ^ j p ¯ j β ^ j
In the case of LD between SNPs, the posterior means can be computed analytically only with an infinitesimal prior. Under a Gaussian infinitesimal prior, the posterior mean effect size is derived as E ( β j | β ^ j ) = D + M N h g 2 I - 1 β ^ j β ^ j ~ N ( D β , D / N ) , f ( β j | β ^ j ) = f ( β j ) e - N 2 ( β ^ - D β ) T D - 1 ( β ^ - D β ) h g 2 β ^ j
The procedure for computing the PRS using LDpred consists of three steps: (1) synchronizing the genotype and summary data, (2) generating LDpred SNP weights, and (3) generating the individual PRS. The first step synchronizes genotype and summary statistics and then generates the coordinated genotype data with the ‘ldpred coord’ command. It requires one genotype file (LD reference) with at least 1,000 individuals of the same ancestry as the individuals for summary statistics. The second step generates an LD information file with a pre-specified LD radius and re-weights the SNP effects with the ‘ldpred gibbs’ command. One LD information file is created with a pre-specified LD radius, but several SNP weight files are generated corresponding to the different values of p (the proportion of causal SNPs). The third step computes the PRS for individuals in the target dataset with the ‘ldpred score’ command. Separate PRS files are generated corresponding to the different values of p. Additionally, LDpred provides a pruning and thresholding option as an alternative method with the ‘ldpred p+t’ command. This option often yields better prediction results than the original LDpred when the sample size of LD reference panel is not large enough.
The construction of a genome-wide PRS using LDpred requires summary statistics from existing large-scale GWAS studies (e.g., the UK Biobank [37-39], DIAGRAM [40]) and an LD reference panel (e.g., the 1000 Genomes project) [41]. A set of candidate PRSs is computed with ranging causal fractions ranging from 0.001 to 1 with p-value thresholding and LD pruning. A range of p-values and pairwise correlations in the LD reference panel are used to include the significantly-associated SNPs for each LD-based clump across the genome with various thresholds [9]. The candidate PRSs are calculated in a validation dataset by multiplying the genotype dosage for each variant by its corresponding weight and summing across all SNPs. The optimal model is selected based on the maximal AUC computed in a validation dataset, and the PRS in the target dataset is then computed. The association between the computed PRS and the target traits is evaluated using linear regression (for a continuous trait) or logistic regression (for a binary trait) with adjustment for covariates (e.g., age, sex, and genotype principal components). The inclusion of such covariates generally leads to more accurate estimates of the PRS and increases the prediction accuracy, but makes it difficult to quantify the exact genetic effects on the target trait. Thus, reporting PRS results with and without important covariates is recommended.
Recently, LDpred-2, a new version of LDpred, was developed to improve predictive performance compared to LDpred [42]. It provides two new options: (1) the ‘sparse’ option, which can make SNP effects exactly 0; and (2) the ‘auto’ option, which learns the tuning parameter p, which is the proportion of causal SNPs, directly from the dataset. LDpred-2 was implemented in the R package ‘bigsnp’.
An alternative to summary-based approaches is to fit the effect sizes of all SNPs simultaneously using BLUP models, which is a more traditional approach for computing the PRS. Fitting all SNPs simultaneously is more appropriate than summary-based approaches, producing more accurate predictors.
GBLUP methods utilize individual-level GWAS data, not summary statistics, to estimate SNP effects using LMMs. The GBLUP model is y=Xβ+g+e, where y is a vector of phenotypes (N×1), X is a matrix of covariates excluding the SNPs (N×C), β is a vector of covariate effects (C×1) and g is a vector of random genetic effects for all individuals with g ~ N ( 0 , σ g 2 A ) e ~ N ( 0 , σ e 2 I ) g ^ = E ( g | y ) = σ g 2 A ( σ g 2 A + σ e 2 I ) - 1 ( y - X β ) u ~ N ( 0 , σ u 2 I ) u ^ = E ( u | g ^ ) = W T A - 1 g ^ / M g ^ g ^ n e w = W n e w u ^ u ^ g ^ n e w
The GBLUP models require individual-level genotype and phenotype data for training, but this is not always possible. Instead, summary SBLUP models can be utilized by approximating individual-level genotype and phenotype data using summary statistics and a reference panel [14]. The SBLUP model is similar to the LDpred model, but it only considers the infinitesimal case, which corresponds to the LDpred-Inf model. The SNP effects (i.e., SNP BLUP) in the RRBLUP model are re-written as u ^ = ( W T W + λ I ) - 1 W T y λ = σ e 2 σ u 2 E ( W T W ) = V T V * ( n t n r ) = B E ( W T y ) = d i a g ( B ) β ^ β ^ u ^ = ( B + λ I ) - 1 d i a g ( B ) β ^ B = V T V * ( n t n r ) h g 2 = M σ u 2 σ p 2 σ p 2 λ = M ( 1 h g 2 - 1 ) g ^ n e w = W n e w u ^
GCTA software was initially designed to estimate SNP-based heritability and has been extended for many other genetic analyses including GBLUP and SBLUP. For GBLUP analysis, the GRM (A) is first estimated from the training genotype data with the ‘--make-grm’ option, and then the individual BLUP (g ^ u ^ g ^ u ^
BMR methods extend the standard LMM by including an alternative prior for SNP effects, further improving prediction accuracy [14,15,17].
The BMR model, BayesR [15,16] assumes that the phenotype is related to set of SNPs under a multiple linear regression model: y=Xβ+e where y is a vector of centered phenotypes (N×1), X is a matrix of standardized genotypes (N×M), β is a vector of SNP effects (M×1) and e is a vector of random errors with e ~ N ( 0 , σ e 2 I ) P ( β j | π , σ β 2 ) = ∑ c = 1 C π c N ( β j | 0 , σ β c 2 ) N ( β j | 0 , σ β c 2 ) σ β c 2 σ β 2 = ( σ β 1 2 , . . . , σ β C 2 ) P ( β | π , σ β 2 , σ e 2 ) ∝ P ( β | π , σ β 2 ) P ( π ) P ( σ β 2 ) P ( σ e 2 ) E ( β | π , σ β 2 , σ e 2 )
The BayesR model with individual-level data was extended to utilize summary statistics from GWAS studies in SBayesR [17]. The SBayesR model relates estimates of multiple regression coefficients (β) to estimates of regression coefficients from M simple linear regression (b) by multiplying y=Xβ+e by D-1XT, where D = diag ( x 1 T x 1 , . . . , x M T x M ) B = D - 1 2 X T X D - 1 2 b = D - 1 2 B D - 1 2 β + D - 1 X T e ( β ) : L ( β ; b , D , B ) : = N ( b ; D - 1 2 B D 1 2 β , D - 1 2 B D - 1 2 σ e 2 ) D ^ = d i a g ( N 1 , . . . , N M ) β ^ P ( β j | π , σ β 2 ) = ∑ c = 1 C π c N ( β j | 0 , γ C σ β 2 ) P ( β | b , D , B , π , σ β 2 , σ e 2 ) ∝ P ( β | D , B , σ β 2 , σ e 2 ) P ( b | β , D , B ) P ( π ) P ( σ β 2 ) P ( σ e 2 ) E ( β | b , D , B , π , σ β 2 , σ e 2 )
GCTB (Genome-wide Complex Trait Bayesian Analysis, https://cnsgenomics.com/software/gctb/) is a software tool that contains a family of Bayesian LMMs for complex trait analyses using GWAS SNPs. First of all, GCTB specifies the Bayesian alphabet for the analysis with the option ‘--bayes': R for BayesR. The options ‘--pi 0.05’ (a starting value for sampling π) and ‘--hsq 0.5’ (a starting value for sampling σ β 2 σ e 2
GBLUP-based methods implicitly assume an infinitesimal genetic architecture, whereas in reality complex traits or diseases are estimated to have roughly only a few thousand causal SNPs in the genome [45,46]. This fact has provided motivation for efforts to construct a PRS that accommodates a non-infinitesimal genetic architecture using penalized regression-based prediction methods.
Penalized regression methods such as the lasso [18,19], the elastic net [20], the adaptive lasso [47], or other statistical learning methods [48] have previously been evaluated for genomic risk prediction [49,50]. The traditional linear regression model is y=Xβ+e, where y is a vector of phenotypic values (N×1), X is a matrix of genotypes (N×M), β is a vector of SNP effects (M×1) and e is random error with e ~ N ( 0 , σ e 2 I ) f ( β ) = ( y - X β ) T ( y - X β ) + λ α β 1 + ( 1 - α β 2 2 / 2 ) | β | 1 = ∑ j = 1 M β j | β | 2 = ∑ j = 1 M β j 2
The lassosum is a method for computing lasso or elastic net estimates using GWAS summary statistics and an LD reference panel [21]. The object function for lasso is given by f ( β ) = ( y - X β ) T ( y - X β ) + λ | β | 1 = y y T - 2 β T X T y + β T X T X β + λ | β | 1 y y T - 2 β T r + β T R β + λ | β | 1 R = ( 1 - s ) X r T X r + s I
The most popular tool for lasso, ridge, and elastic net regression is ‘glmnet’ in R (https://cran.r-project.org/web/packages/glmnet/). The ‘glmnet’ package fits a generalized linear model via the penalized maximum likelihood approach. It is not originally designed for GWAS studies, but it is widely used for PRS analyses due to its computational efficiency. The lassosum is a R package or standalone software for performing lasso or elastic net regression with summary data and an LD reference panel. The reference panel is assumed to be in the PLINK format, and the GWAS summary statistics are given as data.frame in R.
Recent studies have shown that GWAS of related phenotypes further improve the accuracy of polygenic predictions [23,25,51]. Human complex traits and disease traits share genetic architecture with other genetically related traits; therefore, the integration of multiple traits through appropriate methods would achieve improvement in prediction accuracy.
In order to utilize multiple traits to improve prediction accuracy, the RRBLUP and GBLUP methods are extended to the bivariate ridge regression method [52] and MTGBLUP [13,16,43,44], which treat genetic effects as random to obtain individual BLUP and SNP BLUP using one or more genetically correlated traits. The GBLUP models are readily extended to multiple traits (T traits): yi=Xiβi+gi+ei=Xiβi+Wiui+ei (i.e. gi=Wiui) where g i ~ N ( 0 , σ g i 2 A ) e i ~ N ( 0 , σ e i 2 I ) ( g ^ 1 , . . . , g ^ T ) T ( u ^ 1 , . . . , u ^ T ) T
The individual BLUP in a validation sample ( g ^ 1 , n e w , . . . , g ^ T , n e w ) T g ^ 1 , n e w ⋮ g ^ T , n e w = W 1 , n e w ⋯ 0 ⋮ ⋱ ⋮ 0 ⋯ W T , n e w u ^ 1 ⋮ u ^ T u ^ 1 ⋮ u ^ T ( u ^ 1 , . . . , u ^ T ) T
The wMT-SBLUP [22] creates the PRS as a weighted index that combines published GWAS summary statistics across many different traits. The SNP BLUP for T traits can be re-written as u ^ 1 ⋮ u ^ T = W T W + ∑ ϵ ∑ u - 1 ⊗ I M - 1 W T y W = W 1 ⋯ 0 ⋮ ⋱ ⋮ 0 ⋯ W T , ∑ ϵ = σ ϵ 1 2 ⋯ 0 ⋮ ⋱ ⋮ 0 ⋯ σ ϵ T 2 , ∑ u = σ g 1 2 ⋯ σ g 1 T ⋮ ⋱ ⋮ σ g T 1 ⋯ σ g T 2 E ( W i T W i ) = N i L E ( W i T y i ) = N i β ^ i β ^ i u ^ 1 ⋮ u ^ T = I k ⊗ L + ∑ ϵ ∑ u - 1 N - 1 ⊗ I M - 1 β ^ β ^ = β ^ 1 ⋮ β ^ T , N = d i a g ( N 1 , ⋯ , N T )
To utilize multiple traits for PRS computation, the CTPR method was developed [23]. The SNP coefficients are estimated using the following equation: β ^ = a r g m i n R S S ( β ) β + p λ 1 s p ( β ) + p λ 2 c t p ( β ) p λ 1 s p ( β ) p λ 2 c t p ( β ) y ^ t = X t β ^ β ^ y ^ t
Genetic risk prediction in diverse populations currently lags far behind risk prediction in European samples [25,53]. Striking examples include a reported relative decrease of 53%‒89% in schizophrenia risk prediction accuracy in Japanese and African-American populations [12] and 70%‒80% in BMI and type 2 diabetes prediction accuracy in those of African ancestry [54] compared to Europeans in studies using European training data due to between-population differences in population allele frequencies and patterns of LD. An alternative is to use training data from the target population, but this generally implies a much lower training sample size, reducing prediction accuracy. A recent method that incorporates training data from European and non-European populations improves prediction accuracy by using XP-BLUP [24] with the use of European-discovered SNPs and population-specific weights or by using a multi-ethnic PRS [25] and multi-ancestry PRS [26] with averages across all admixed individuals.
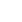
We have reviewed statistical models and computational tools for PRS computation. We have demonstrated a variety of statistical models for genomic risk prediction using individual-level data and/or summary statistics and showed how to improve prediction accuracy with multiple traits and multiple populations. Furthermore, we have introduced recent computational tools to conduct PRS analyses based on the statistical models, and explained how to specify the parameters and how to execute the software in detail. We also summarized which statistical models and software are best for specific situations based on data type (GWAS summary statistics or individual-level GWAS data), sample size, the LD reference panel, the number of traits, and the number of ethnicities, as shown in Fig. 1. The summary-based PRS methods such as PLINK, LDpred, and SBLUP offer advantages in computational cost over PRS methods using individual-level data such as GBLUP and the lasso method. This is because the computation time of summary-based PRS methods does not increase with the number of individuals in the study. This advantage has motivated the recent development of various summary-based methods in conjunction with LD information, although PRS methods using individual-level data could generate more accurate PRS. With recent large-sample GWAS data, summary-based methods are generally utilized due to their computational efficiencies, while PRS methods using individual-level data are still usable for computing more accurate PRS.
Despite the existence of various PRS methods, there are some areas in which further research on PRS is required. To improve prediction accuracy, we need novel statistical models and software that leverage information from multiple disease outcomes and multiple ethnicities based on individual-level genotype data and/or summary statistics from large-scale biobanks. It is also necessary to develop methods with the ability to predict diverse disease traits, such as cardiovascular disease and type 2 diabetes, with sufficient accuracy (to the extent allowable by disease heritability), and then these models need to be extended to utilize multiple ethnicities by incorporating information on LD to further improve prediction accuracy.
Moreover, with advances in high-throughput molecular assays (e.g., RNA-seq and ChIP-seq), it has been shown that disease risk SNPs are enriched in a broad array of functional regions, including regulatory features that are often tissue-specific, providing a novel source of information for improved prediction accuracy. It has been further shown that these molecular features can be predicted from genetic variants, enabling the prediction of gene expression in GWAS cohorts to perform transcriptome-wide association studies and to identify putative susceptibility genes. The accurate prediction of individual molecular features is now an emerging tool for discovering novel disease loci and characterizing biological mechanisms at the thousands of GWAS loci that have already been published. Data collection efforts of an unprecedented scale are now being seen in the areas of functional genomics and disease genetics. Such datasets can help to prioritize causal features and further improve prediction accuracy.
We conclude by emphasizing the importance of creating accurate PRS for a wide range of complex traits and diseases. The PRS provides an estimate of genetic predisposition (also called genetic susceptibility) for a complex trait or disease at the individual level, which refers to the likelihood of developing a particular trait or disease based on a genotype profile. The goal of PRS analysis is to identify individuals at an elevated risk of diseases on the basis of genetic variants in combination with clinical covariates. Therefore, the more accurate PRS we obtain, the better we can identify disease risk and the better we can provide treatment and prevention strategies. Personalized medicine based on accurate PRS will have a considerable impact on the treatment process and quality of life in the near future.
Acknowledgments
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (NRF-2020R1C1C1A01012657) and Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2021R1A6A1A10044154). This work was supported by Soongsil University Research Fund.
Fig. 1.
Best statistical models and software based on data type, sample size, LD reference panel, and the number of traits and ethnicities. CTPR, cross-trait penalized regression; GBLUP, genomic BLUP; GWAS, genome-wide association studies; LD, linkage disequilibrium; MTGBLUP, multi-trait GBLUP; SBLUP, statistics BLUP; wMT-SBLUP, weighted multi-trait SBLUP.
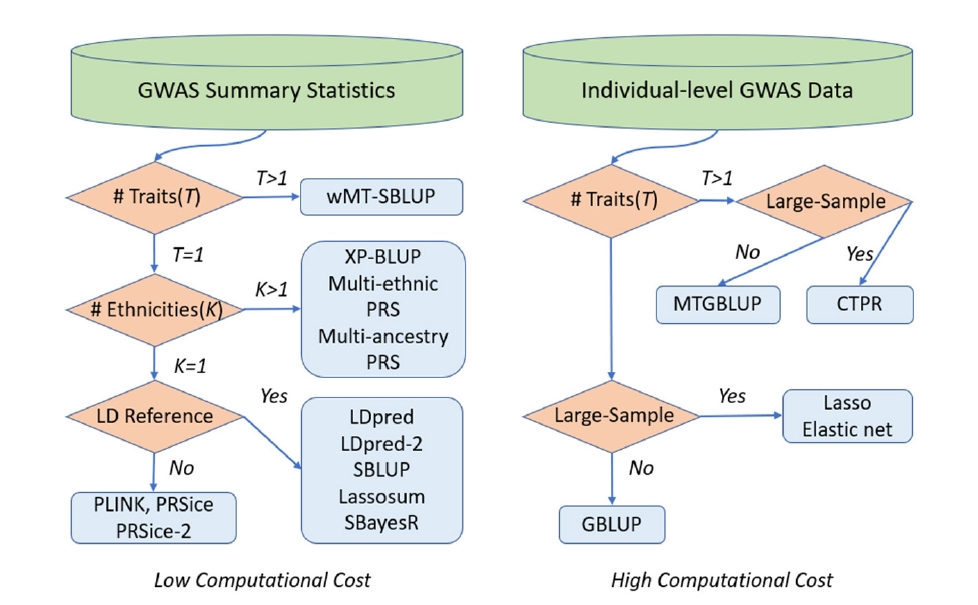
Table 1.
List of PRS methods, underlying statistical models, computational tools, and required data
References
1. Visscher PM, Wray NR, Zhang Q, Sklar P, McCarthy MI, Brown MA, et al. 10 Years of GWAS discovery: biology, function, and translation. Am J Hum Genet 2017;101:5–22.



2. Visscher PM, Brown MA, McCarthy MI, Yang J. Five years of GWAS discovery. Am J Hum Genet 2012;90:7–24.



3. Chatterjee N, Shi J, Garcia-Closas M. Developing and evaluating polygenic risk prediction models for stratified disease prevention. Nat Rev Genet 2016;17:392–406.



4. Yang J, Zeng J, Goddard ME, Wray NR, Visscher PM. Concepts, estimation and interpretation of SNP-based heritability. Nat Genet 2017;49:1304–1310.


5. Wang K, Gaitsch H, Poon H, Cox NJ, Rzhetsky A. Classification of common human diseases derived from shared genetic and environmental determinants. Nat Genet 2017;49:1319–1325.



6. Munoz M, Pong-Wong R, Canela-Xandri O, Rawlik K, Haley CS, Tenesa A. Evaluating the contribution of genetics and familial shared environment to common disease using the UK Biobank. Nat Genet 2016;48:980–983.



7. Wray NR, Yang J, Hayes BJ, Price AL, Goddard ME, Visscher PM. Pitfalls of predicting complex traits from SNPs. Nat Rev Genet 2013;14:507–515.



8. Chung W, Zou F. Mixed-effects models for GAW18 longitudinal blood pressure data. BMC Proc 2014;8(Suppl 1):S87.



9. Khera AV, Chaffin M, Aragam KG, Haas ME, Roselli C, Choi SH, et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat Genet 2018;50:1219–1224.



10. Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 2007;81:559–575.



11. Yang J, Lee SH, Goddard ME, Visscher PM. GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet 2011;88:76–82.



12. Vilhjalmsson BJ, Yang J, Finucane HK, Gusev A, Lindstrom S, Ripke S, et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am J Hum Genet 2015;97:576–592.



13. de Los Campos G, Vazquez AI, Fernando R, Klimentidis YC, Sorensen D. Prediction of complex human traits using the genomic best linear unbiased predictor. PLoS Genet 2013;9:e1003608.



14. Robinson MR, Kleinman A, Graff M, Vinkhuyzen AA, Couper D, Miller MB, et al. Genetic evidence of assortative mating in humans. Nature Human Behaviour 2017;1:0016.

15. Erbe M, Hayes BJ, Matukumalli LK, Goswami S, Bowman PJ, Reich CM, et al. Improving accuracy of genomic predictions within and between dairy cattle breeds with imputed high-density single nucleotide polymorphism panels. J Dairy Sci 2012;95:4114–4129.


16. Maier R, Moser G, Chen GB, Ripke S, Cross-Disorder Working Group of the Psychiatric Genomics Consortium, Coryell W, et al. Joint analysis of psychiatric disorders increases accuracy of risk prediction for schizophrenia, bipolar disorder, and major depressive disorder. Am J Hum Genet 2015;96:283–294.


17. Lloyd-Jones LR, Zeng J, Sidorenko J, Yengo L, Moser G, Kemper KE, et al. Improved polygenic prediction by Bayesian multiple regression on summary statistics. Nat Commun 2019;10:5086.



18. Tibshirani R. Regression shrinkage and selection via the Lasso. J R Stat Soc Series B Stat Methodol 1996;58:267–288.

19. Lello L, Avery SG, Tellier L, Vazquez AI, de Los Campos G, Hsu SD. Accurate genomic prediction of human height. Genetics 2018;210:477–497.



20. Zou H, Hastie T. Regularization and variable selection via the elastic net. J R Stat Soc Series B Stat Methodol 2005;67:301–320.

21. Mak TS, Porsch RM, Choi SW, Zhou X, Sham PC. Polygenic scores via penalized regression on summary statistics. Genet Epidemiol 2017;41:469–480.


22. Maier RM, Zhu Z, Lee SH, Trzaskowski M, Ruderfer DM, Stahl EA, et al. Improving genetic prediction by leveraging genetic correlations among human diseases and traits. Nat Commun 2018;9:989.



23. Chung W, Chen J, Turman C, Lindstrom S, Zhu Z, Loh PR, et al. Efficient cross-trait penalized regression increases prediction accuracy in large cohorts using secondary phenotypes. Nat Commun 2019;10:569.



24. Coram MA, Fang H, Candille SI, Assimes TL, Tang H. Leveraging multi-ethnic evidence for risk assessment of quantitative traits in minority populations. Am J Hum Genet 2017;101:218–226.



25. Marquez-Luna C, Loh PR, South Asian Type 2 Diabetes (SAT2D) Consortium, SIGMA Type 2 Diabetes Consortium, Price AL. Multiethnic polygenic risk scores improve risk prediction in diverse populations. Genet Epidemiol 2017;41:811–823.



26. Cavazos TB, Witte JS. Inclusion of variants discovered from diverse populations improves polygenic risk score transferability. HGG Adv 2021;2:100017.


27. International Schizophrenia Consortium, Purcell SM, Wray NR, Stone JL, Visscher PM, O'Donovan MC, et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 2009;460:748–752.



28. Pasaniuc B, Price AL. Dissecting the genetics of complex traits using summary association statistics. Nat Rev Genet 2017;18:117–127.


29. Wray NR, Yang J, Goddard ME, Visscher PM. The genetic interpretation of area under the ROC curve in genomic profiling. PLoS Genet 2010;6:e1000864.



30. Lee SH, Goddard ME, Wray NR, Visscher PM. A better coefficient of determination for genetic profile analysis. Genet Epidemiol 2012;36:214–224.


31. Daetwyler HD, Villanueva B, Woolliams JA. Accuracy of predicting the genetic risk of disease using a genome-wide approach. PLoS One 2008;3:e3395.



32. Visscher PM, Yang J, Goddard ME. A commentary on 'common SNPs explain a large proportion of the heritability for human height' by Yang et al. (2010). Twin Res Hum Genet 2010;13:517–524.


33. Locke AE, Kahali B, Berndt SI, Justice AE, Pers TH, Day FR, et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 2015;518:197–206.


34. Polychronakos C, Li Q. Understanding type 1 diabetes through genetics: advances and prospects. Nat Rev Genet 2011;12:781–792.


35. Euesden J, Lewis CM, O'Reilly PF. PRSice: polygenic risk score software. Bioinformatics 2015;31:1466–1468.


36. Choi SW, O'Reilly PF. PRSice-2: Polygenic Risk Score software for biobank-scale data. Gigascience 2019;8:giz082.



37. Allen NE, Sudlow C, Peakman T, Collins R, Biobank UK. UK biobank data: come and get it. Sci Transl Med 2014;6:224e. d224.

38. UKBiobank. Genotyping and Quality Control of UK Biobank, a Large-Scale, Extensively Phenotyped Prospective Resource. Cheshire: UK Biobank, 2015.
39. UKBiobank. UK Biobank: Genotyping and Imputation Data Release. Cheshire: UK Biobank, 2018.
40. Morris AP, Voight BF, Teslovich TM, Ferreira T, Segre AV, Steinthorsdottir V, et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat Genet 2012;44:981–990.



41. Mahajan A, Taliun D, Thurner M, Robertson NR, Torres JM, Rayner NW, et al. Fine-mapping type 2 diabetes loci to single-variant resolution using high-density imputation and islet-specific epigenome maps. Nat Genet 2018;50:1505–1513.



42. Prive F, Arbel J, Vilhjalmsson BJ. LDpred2: better, faster, stronger. Bioinformatics 2020;36:5424–5431.



43. Meuwissen TH, Hayes BJ, Goddard ME. Prediction of total genetic value using genome-wide dense marker maps. Genetics 2001;157:1819–1829.



44. Habier D, Fernando RL, Dekkers JC. The impact of genetic relationship information on genome-assisted breeding values. Genetics 2007;177:2389–2397.



45. Loh PR, Tucker G, Bulik-Sullivan BK, Vilhjalmsson BJ, Finucane HK, Salem RM, et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat Genet 2015;47:284–290.



46. Stahl EA, Wegmann D, Trynka G, Gutierrez-Achury J, Do R, Voight BF, et al. Bayesian inference analyses of the polygenic architecture of rheumatoid arthritis. Nat Genet 2012;44:483–489.



48. Abraham G, Kowalczyk A, Zobel J, Inouye M. Performance and robustness of penalized and unpenalized methods for genetic prediction of complex human disease. Genet Epidemiol 2013;37:184–195.


49. Abraham G, Tye-Din JA, Bhalala OG, Kowalczyk A, Zobel J, Inouye M. Accurate and robust genomic prediction of celiac disease using statistical learning. PLoS Genet 2014;10:e1004137.



50. Wu TT, Chen YF, Hastie T, Sobel E, Lange K. Genome-wide association analysis by lasso penalized logistic regression. Bioinformatics 2009;25:714–721.



51. Turley P, Walters RK, Maghzian O, Okbay A, Lee JJ, Fontana MA, et al. Multi-trait analysis of genome-wide association summary statistics using MTAG. Nat Genet 2018;50:229–237.



52. Li C, Yang C, Gelernter J, Zhao H. Improving genetic risk prediction by leveraging pleiotropy. Hum Genet 2014;133:639–650.


- TOOLS
-
METRICS

-
- 7 Crossref
- 0 Scopus
- 5,099 View
- 244 Download
- Related articles in GNI








